Documentation (Latest | Stable) | DGL at a glance | Model Tutorials | Discussion Forum
DGL is an easy-to-use, high performance and scalable Python package for deep learning on graphs. DGL is framework agnostic, meaning if a deep graph model is a component of an end-to-end application, the rest of the logics can be implemented in any major frameworks, such as PyTorch, Apache MXNet or TensorFlow.

Figure: DGL Overall Architecture
06/11/2020: Amazon Shanghai AI Lab and AWS Deep Engine Science team working along with academic collaborators from the University of Minnesota, The Ohio State University, and Hunan University have created the Drug Repurposing Knowledge Graph (DRKG) and a set of machine learning tools, DGL-KE, that can be used to prioritize drugs for repurposing studies. DRKG is a comprehensive biological knowledge graph that relates human genes, compounds, biological processes, drug side effects, diseases and symptoms. DRKG includes, curates, and normalizes information from six publicly available databases and data that were collected from recent publications related to Covid-19. It has 97,238 entities belonging to 13 types of entities, and 5,874,261 triplets belonging to 107 types of relations. More about the dataset is in this blogpost.
03/31/2020: The new v0.4.3 release includes official TensorFlow support, with 15 popular GNN modules. DGL-KE and DGL-LifeSci, two packages for knowledge graph embedding and chemi- and bio-informatics respectively, have graduated as standalone packages and can be installed by pip and conda. The new release provides full support of graph sampling on heterogeneous graphs, with multi-GPU acceleration. See our new feature walkthrough and release note.
A data scientist may want to apply a pre-trained model to your data right away. For this you can use DGL's Application packages, formally Model Zoo. Application packages are developed for domain applications, as is the case for DGL-LifeScience. We will soon add model zoo for knowledge graph embedding learning and recommender systems. Here's how you will use a pretrained model:
from dgllife.data import Tox21
from dgllife.model import load_pretrained
from dgllife.utils import smiles_to_bigraph, CanonicalAtomFeaturizer
dataset = Tox21(smiles_to_bigraph, CanonicalAtomFeaturizer())
model = load_pretrained('GCN_Tox21') # Pretrained model loaded
model.eval()
smiles, g, label, mask = dataset[0]
feats = g.ndata.pop('h')
label_pred = model(g, feats)
print(smiles) # CCOc1ccc2nc(S(N)(=O)=O)sc2c1
print(label_pred[:, mask != 0]) # Mask non-existing labels
# tensor([[ 1.4190, -0.1820, 1.2974, 1.4416, 0.6914,
# 2.0957, 0.5919, 0.7715, 1.7273, 0.2070]])Further reading: DGL is released as a managed service on AWS SageMaker, see the medium posts for an easy trip to DGL on SageMaker(part1 and part2).
Researchers can start from the growing list of models implemented in DGL. Developing new models does not mean that you have to start from scratch. Instead, you can reuse many pre-built modules. Here is how to get a standard two-layer graph convolutional model with a pre-built GraphConv module:
from dgl.nn.pytorch import GraphConv
import torch.nn.functional as F
# build a two-layer GCN with ReLU as the activation in between
class GCN(nn.Module):
def __init__(self, in_feats, h_feats, num_classes):
super(GCN, self).__init__()
self.gcn_layer1 = GraphConv(in_feats, h_feats)
self.gcn_layer2 = GraphConv(h_feats, num_classes)
def forward(self, graph, inputs):
h = self.gcn_layer1(graph, inputs)
h = F.relu(h)
h = self.gcn_layer2(graph, h)
return hNext level down, you may want to innovate your own module. DGL offers a succinct message-passing interface (see tutorial here). Here is how Graph Attention Network (GAT) is implemented (complete codes). Of course, you can also find GAT as a module GATConv:
import torch.nn as nn
import torch.nn.functional as F
# Define a GAT layer
class GATLayer(nn.Module):
def __init__(self, in_feats, out_feats):
super(GATLayer, self).__init__()
self.linear_func = nn.Linear(in_feats, out_feats, bias=False)
self.attention_func = nn.Linear(2 * out_feats, 1, bias=False)
def edge_attention(self, edges):
concat_z = torch.cat([edges.src['z'], edges.dst['z']], dim=1)
src_e = self.attention_func(concat_z)
src_e = F.leaky_relu(src_e)
return {'e': src_e}
def message_func(self, edges):
return {'z': edges.src['z'], 'e':edges.data['e']}
def reduce_func(self, nodes):
a = F.softmax(nodes.mailbox['e'], dim=1)
h = torch.sum(a * nodes.mailbox['z'], dim=1)
return {'h': h}
def forward(self, graph, h):
z = self.linear_func(h)
graph.ndata['z'] = z
graph.apply_edges(self.edge_attention)
graph.update_all(self.message_func, self.reduce_func)
return graph.ndata.pop('h')Microbenchmark on speed and memory usage: While leaving tensor and autograd functions to backend frameworks (e.g. PyTorch, MXNet, and TensorFlow), DGL aggressively optimizes storage and computation with its own kernels. Here's a comparison to another popular package -- PyTorch Geometric (PyG). The short story is that raw speed is similar, but DGL has much better memory management.
| Dataset | Model | Accuracy | Time PyG DGL |
Memory PyG DGL |
|---|---|---|---|---|
| Cora | GCN GAT |
81.31 ± 0.88 83.98 ± 0.52 |
0.478 0.666 1.608 1.399 |
1.1 1.1 1.2 1.1 |
| CiteSeer | GCN GAT |
70.98 ± 0.68 69.96 ± 0.53 |
0.490 0.674 1.606 1.399 |
1.1 1.1 1.3 1.1 |
| PubMed | GCN GAT |
79.00 ± 0.41 77.65 ± 0.32 |
0.491 0.690 1.946 1.393 |
1.1 1.1 1.6 1.1 |
| GCN | 93.46 ± 0.06 | OOM 28.6 | OOM 11.7 | |
| Reddit-S | GCN | N/A | 29.12 9.44 | 15.7 3.6 |
Table: Training time(in seconds) for 200 epochs and memory consumption(GB)
Here is another comparison of DGL on TensorFlow backend with other TF-based GNN tools (training time in seconds for one epoch):
| Dateset | Model | DGL | GraphNet | tf_geometric |
|---|---|---|---|---|
| Core | GCN | 0.0148 | 0.0152 | 0.0192 |
| GCN | 0.1095 | OOM | OOM | |
| PubMed | GCN | 0.0156 | 0.0553 | 0.0185 |
| PPI | GCN | 0.09 | 0.16 | 0.21 |
| Cora | GAT | 0.0442 | n/a | 0.058 |
| PPI | GAT | 0.398 | n/a | 0.752 |
High memory utilization allows DGL to push the limit of single-GPU performance, as seen in below images.
 |
 |
|---|
Scalability: DGL has fully leveraged multiple GPUs in both one machine and clusters for increasing training speed, and has better performance than alternatives, as seen in below images.
 |
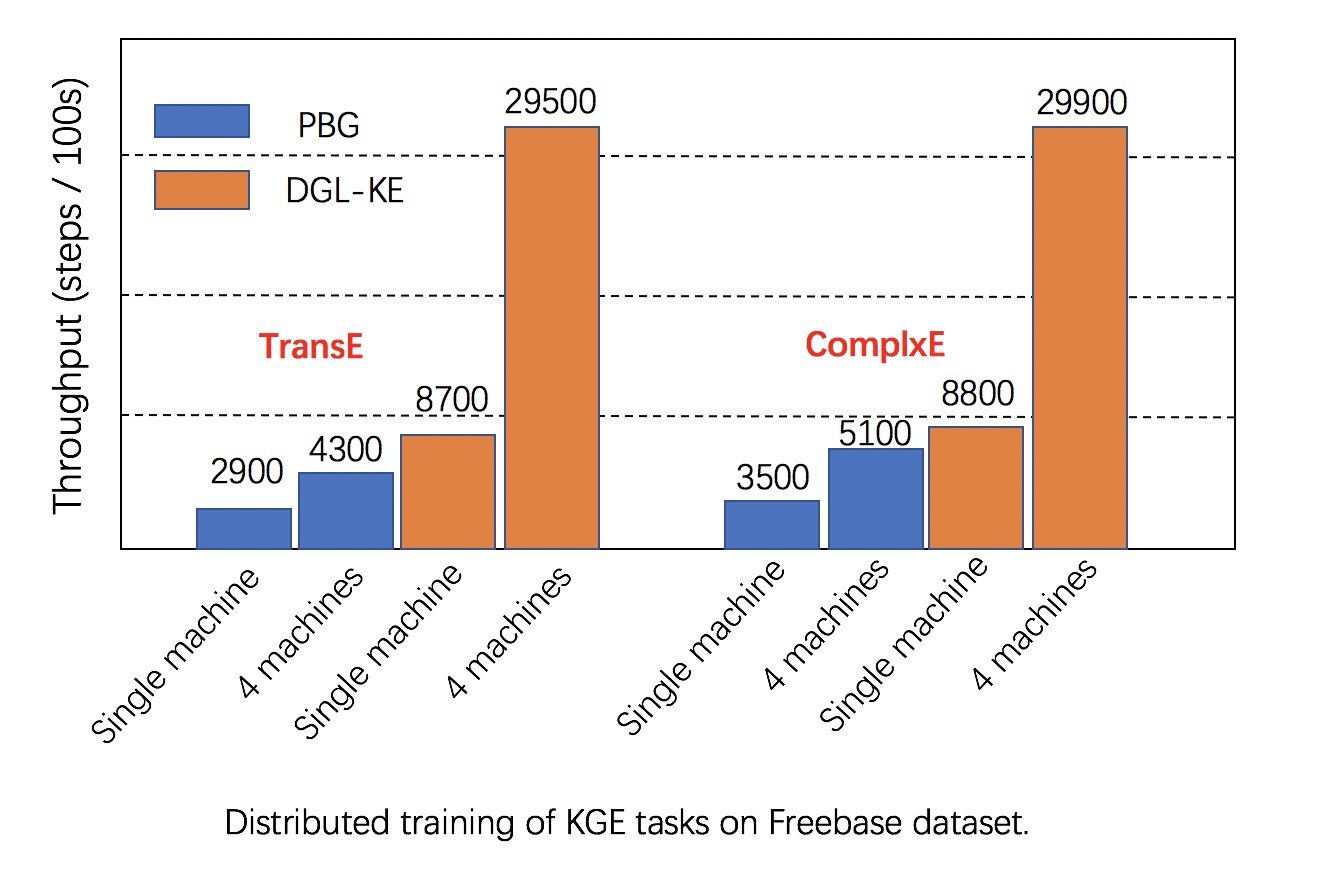 |
|---|
Further reading: Detailed comparison of DGL and other Graph alternatives can be found here.
Overall there are 30+ models implemented by using DGL:
- DGL-LifeSci, previously DGL-Chem
- DGL-KE
- DGL-RecSys(coming soon)
We are currently in Beta stage. More features and improvements are coming.
DGL should work on
- all Linux distributions no earlier than Ubuntu 16.04
- macOS X
- Windows 10
DGL requires Python 3.5 or later.
Right now, DGL works on PyTorch 1.2.0+, MXNet 1.5.1+, and TensorFlow 2.1.0+.
conda install -c dglteam dgl # cpu version
conda install -c dglteam dgl-cuda9.0 # CUDA 9.0
conda install -c dglteam dgl-cuda9.2 # CUDA 9.2
conda install -c dglteam dgl-cuda10.0 # CUDA 10.0
conda install -c dglteam dgl-cuda10.1 # CUDA 10.1
conda install -c dglteam dgl-cuda10.2 # CUDA 10.2
| Latest Nightly Build Version | Stable Version | |
|---|---|---|
| CPU | pip install --pre dgl |
pip install dgl |
| CUDA 9.0 | pip install --pre dgl-cu90 |
pip install dgl-cu90 |
| CUDA 9.2 | pip install --pre dgl-cu92 |
pip install dgl-cu92 |
| CUDA 10.0 | pip install --pre dgl-cu100 |
pip install dgl-cu100 |
| CUDA 10.1 | pip install --pre dgl-cu101 |
pip install dgl-cu101 |
| CUDA 10.2 | pip install --pre dgl-cu102 |
pip install dgl-cu102 |
Refer to the guide here.
| Releases | Date | Features |
|---|---|---|
| v0.4.3 | 03/31/2020 | - TensorFlow support - DGL-KE - DGL-LifeSci - Heterograph sampling APIs (experimental) |
| v0.4.2 | 01/24/2020 | - Heterograph support - TensorFlow support (experimental) - MXNet GNN modules |
| v0.3.1 | 08/23/2019 | - APIs for GNN modules - Model zoo (DGL-Chem) - New installation |
| v0.2 | 03/09/2019 | - Graph sampling APIs - Speed improvement |
| v0.1 | 12/07/2018 | - Basic DGL APIs - PyTorch and MXNet support - GNN model examples and tutorials |
Check out the open source book Dive into Deep Learning.
For those who are new to graph neural network, please see the basic of DGL.
For audience who are looking for more advanced, realistic, and end-to-end examples, please see model tutorials.
Please let us know if you encounter a bug or have any suggestions by filing an issue.
We welcome all contributions from bug fixes to new features and extensions.
We expect all contributions discussed in the issue tracker and going through PRs. Please refer to our contribution guide.
If you use DGL in a scientific publication, we would appreciate citations to the following paper:
@article{wang2019dgl,
title={Deep Graph Library: Towards Efficient and Scalable Deep Learning on Graphs},
url={https://arxiv.org/abs/1909.01315},
author={Wang, Minjie and Yu, Lingfan and Zheng, Da and Gan, Quan and Gai, Yu and Ye, Zihao and Li, Mufei and Zhou, Jinjing and Huang, Qi and Ma, Chao and Huang, Ziyue and Guo, Qipeng and Zhang, Hao and Lin, Haibin and Zhao, Junbo and Li, Jinyang and Smola, Alexander J and Zhang, Zheng},
journal={ICLR Workshop on Representation Learning on Graphs and Manifolds},
year={2019}
}
DGL is developed and maintained by NYU, NYU Shanghai, AWS Shanghai AI Lab, and AWS MXNet Science Team.
DGL uses Apache License 2.0.



